 
However these products are no longer under development. QIAGEN Ingenuity Pathway Analysis QIAGEN OmicSoft Suite ‘Omics Databases. Hardware Įarly on, the company initially presented own-developed high-performance computing solutions, focusing on accelerating open source algorithms such as HMMER, Smith-Waterman and ClustalW, using FPGA technology. QIAGEN CLC Workbench Premium QIAGEN CLC Genomics Workbench QIAGEN CLC Genome Finishing Module QIAGEN CLC Microbial Genomics Module QIAGEN CLC Main Workbench QIAGEN CLC Module and Plugin Overview Interpretation and Visualization. In 2020, CLC bio released a free plug-in that enables workflow execution on AWS directly from the CLC Genomics Workbench desktop software. In 2019, this platform was adapted for and approved for use in AWS GovCloud. In 2017, CLC bio launched their CLC Genomics Cloud Engine as a command-line driven platform for cloud-based bioinformatics workflow execution on Amazon Web Services. 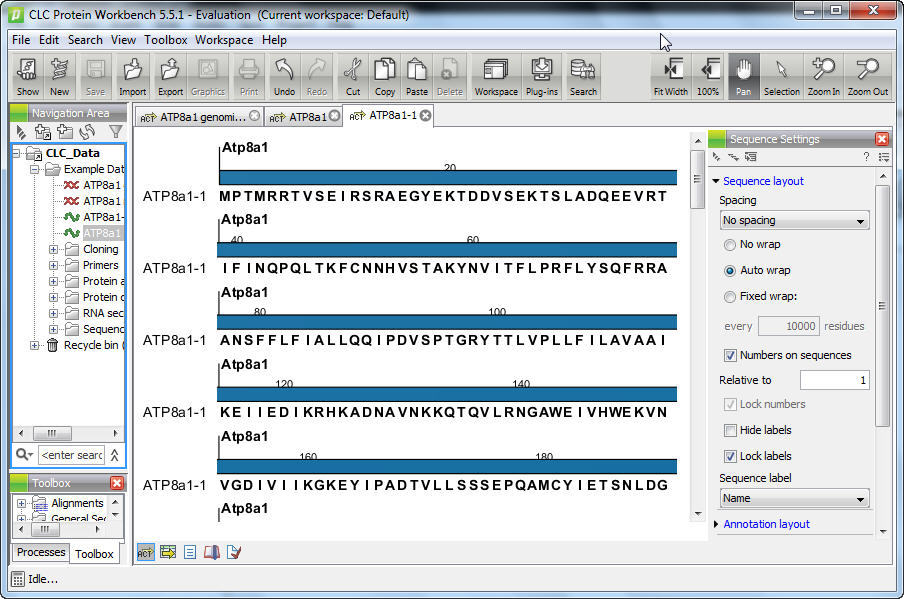 
Features include read mapping and de novo assembly of high-throughput sequencing data, whole-genome detection of SNPs and structural variations, ChIP-seq, RNA-Seq, small RNA analysis, genome finishing, microbial genomics, structural biology, and functions to analyze, visualize, and compare genomic, transcriptomic, and epigenomic data. pathway analysis, genomics, and other omics). Īs additional capabilities were added to the software platform, it was eventually split into several themed Workbenches and plugins with collections of features relevant to different applications (e.g. In 2010, CLC bio was notable as the first commercial platform for bioinformatics analysis that utilized a graphical user interface for building, managing, and deploying analysis workflows as well as command-line tools, a SOAP and REST API, and later, the ability to run containerized tools. CLC bio developed some of their own open source algorithms, as well as their own SIMD-accelerated implementations of several existing popular applications. Software ĬLC bio's main activities were in software development for desktop ( Mac OS X, Windows, and Linux), enterprise, and cloud software for analysis of biological data. ĬLC bio was acquired by QIAGEN in 2013 and merged into its bioinformatics research and development division with several other purchased platforms in 2014. By 2012, it had additional offices in Cambridge, Massachusetts, Tokyo, Taipei and Delhi, with staff largely from research backgrounds (30% having a PhD) and had built a userbase of around 250,000 users in both academic institutions and biotechnology companies. Its product's development was also partly funded by collaborating with researchers on grant-funded projects. But i'm really ignorant about that.Is anybody confident with this kind of analysis? Does anybody have any suggestion? I've a LOT of data and i can't analyze it.CLC bio was a bioinformatics software company that developed a software suite subsequently purchased by QIAGEN.ĬLC bio started commercial activities on Januheadquartered in Aarhus, Denmark. I guess there is a problem with the GFF file. Ant i can't see any value in all the other column of the RNA-seq analyzed sample table. I point out this becouse, when i make the RNA-seq analysis, despite i can see the reads alligning on the annotations, i can't get any expression value for my genes. The only table i can see show me a list of all different chromosomes on my FASTA file. 
The problem is that when i try to look at the annotation table i can't do it. It seems to work, as i can see the annotations on the reference genome in the view window. To annotate the reference genome i use the tool "annotate with a GFF/GTF file". The reference genome is a FASTA file, while annotations are in a GFF format. The CLC Main Workbench was used for sequence analyses, including construction of phylogenetic trees using the MSAs and Neighbor Joining method. To allign all my reads against the 8X reference genome the first think i've to do is to upload my genome on clc and to annotate it. I never used clc before and i'm not a bioinformatic, so i'm sorry if i'm gonna be a bit generic and unprecise. Yet a lack of well integrated analytics for microbial genomics leaves researchers and organisations with the burden of integrating and maintaining all the required bioinformatics, statistics and visualisation tools required to power their research. Its wide variety of features are presented in an intuitive graphical user-interface, for which advanced computer skills are not required. Ever-growing sample volumes demand efficient bioinformatics. QIAGEN CLC Main Workbench is used by tens of thousands of researchers all over the world for DNA, RNA and protein sequence data analysis. Actually i'm trying to look at the transcriptome expression of several semples of treated grapevine leaves against untreated using a 36bp paired ends approach with illumina technology. Differential Expression Analysis: CLC Genomics Workbench Version 20.0.4. From data to discovery with QIAGEN CLC Genomics Workbench Premium. I don't know if anybody is confdent with RNA-seq analysis by clc genomic workbench. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
